Rebecca McGillivary
Quantitative Cell Biologist
Quantitative Cell Biologist


I'm a cell biologist who loves building tools and creating content that helps other researchers succeed. On the technical side, I've developed experimental methods for non-model organisms, written open-source Python tools for data analysis, and collaborated with teams of engineers and computational scientists on imaging technology. On the communications side, I've presented research at international conferences, created interactive museum demonstrations, designed scientific graphics, and authored peer-reviewed publications.
I'm seeking research or technical roles that leverage my ability to bridge experimental biology and computational approaches - whether in AI-accelerated scientific discovery, developer relations, or scientific program management. I am excited about opportunities to combine my deep biology expertise with my passion for technical communication, tool development, and community building. Based in Sunnyvale, CA.
Hartwell Postdoctoral Fellow
University of California, Davis - Starr-Luxton Lab
I was a postdoc in the joint lab of Dr. Daniel Starr and Dr. G.W. Gant Luxton at UC Davis. My goal was to understand how the assembly of the linker of nucleoskeleton and cytoskeleton (LINC) complex affects the mechanical properties of cells, and how the assembly of the LINC complex is regulated. To study how the LINC complex regulates the mechanical properties of cells, I performed high-speed live imaging and microrheology experiments in hTERT-immortalized human airway smooth muscle cells. I also took a biochemical approach to studying LINC complex assembly and stability by utilizing mass photometry and differential scanning fluorimetry.
Graduate Research
University of California, San Francisco - Marshall Lab
For my graduate studies, I joined the lab of Dr. Wallace Marshall at UCSF to study the giant ciliate, Stentor coeruleus. During regeneration, Stentor’s nucleus changes shape in an extreme and developmentally regulated way, making it a powerful new model system to address this fundamental question in cell biology. I discovered the first molecular driver of this process, the nuclear transport factor, CSE1. I utilized immunofluorescence, live-cell imaging, and computational image analysis - often adapting these methods to work with a non-conventional model system.
Undergraduate Research
University of California, Los Angeles - Quinlan Lab
As an undergraduate, I worked with purified proteins to learn how the actin cytoskeleton is organized by actin binding proteins. This project was my introduction into thinking about how biology and physics intersect.
Social media links:
Twitter
LinkedIn
You can also send me an email here:
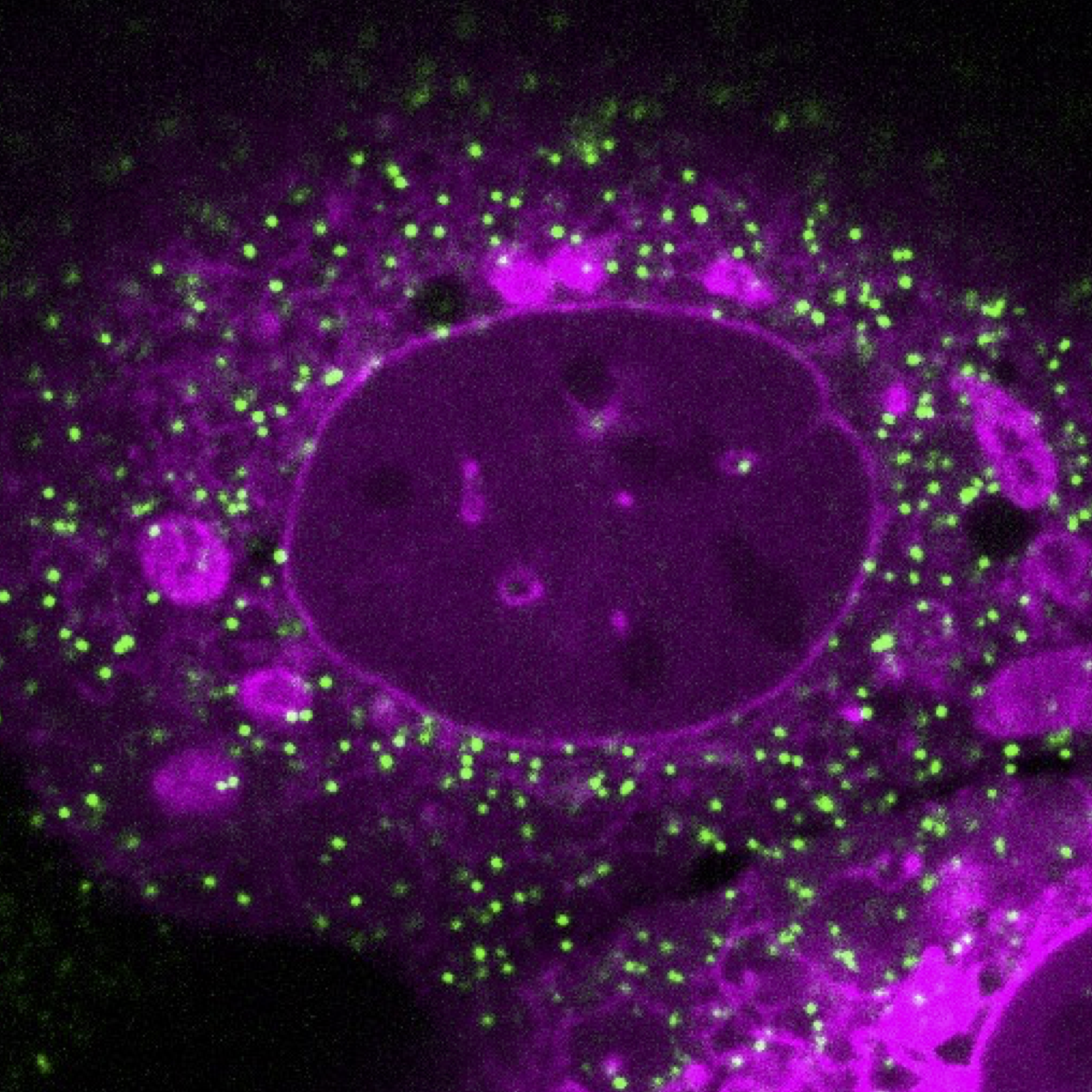
Investigating LINC complex assembly and its effects on cellular mechanics
During my postdoc at UC Davis, I studied the linker of nucleoskeleton and cytoskeleton (LINC) complex. This is a molecular bridge composed of SUN and KASH proteins that span the two membranes of the nuclear envelope, mechanically integrating the nucleus with the cytoskeleton. How do the different LINC complex components affect the mechanical properties of the cell? This is especially important for the biology of smooth muscle, as LINC complexes help smooth muscle adapt to different mechanical environments. To address this question I performed passive microrheology experiments by live imaging human airway smooth muscle cells expressing genetically encoded multimeric nanoparticles (GEMs). I collaborated with a computational scientist in the lab to perform single-particle tracking of the GEMs and calculate their diffusion coefficients. By combining these microrheology experiments with knockdowns of LINC complex proteins, I quantified how the LINC complex affects macromolecular crowding in smooth muscle.
In addition to my cell biology work, I also used a biochemical approach to investigate how the assembly of the LINC complex is regulated. I developed an in vitro assay using mass photometry to quantify SUN protein oligomerization and to determine how this is affected by buffer conditions and mutations. I also utilized differential scanning fluorimetry to measure how buffer conditions affect the stability of SUN proteins. These assays allowed me to investigate LINC assembly in finer detail than has been previously possible.
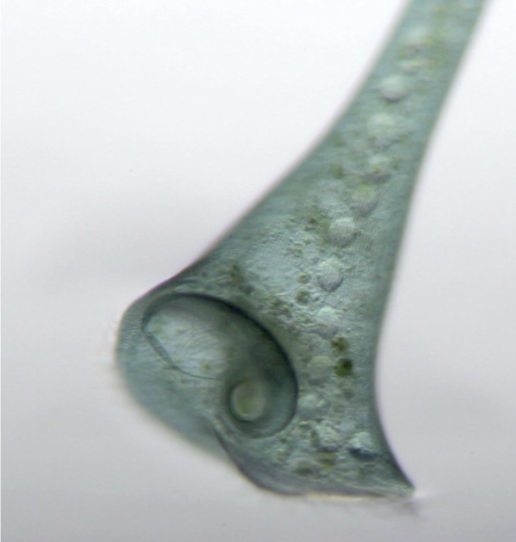

Identified the first molecular driver of the macronuclear shape change cycle in Stentor coeruleus.
During my graduate work in the Marshall Lab at UCSF, I studied Stentor coeruleus, a non-model organism that undergoes a dramatic and developmentally regulated nuclear shape change. During regeneration of Stentor’s oral apparatus, the macronucleus rapidly changes shape: it initially resembles a string of beads, then coalesces into a sphere, and then elongates back into its original moniliform shape. This process provides an excellent opportunity to study how cells regulate the size and shape of their nuclei. I found that the nuclear transport factor, CSE1, plays a role in altering the dynamics of this shape change: CSE1 is needed for the average volume per node to rapidly increase during condensation, and it is needed for the rapid change in the median number of nodes per macronucleus. CSE1 is also necessary for Stentor to maintain the moniliform shape of its macronucleus after regeneration is completed. Our model is that the transport of proteins into the nucleus alters the volume of the macronucleus, which in turn, alters the shape of the macronucleus due to constraints on the surface area of the macronuclear envelope.
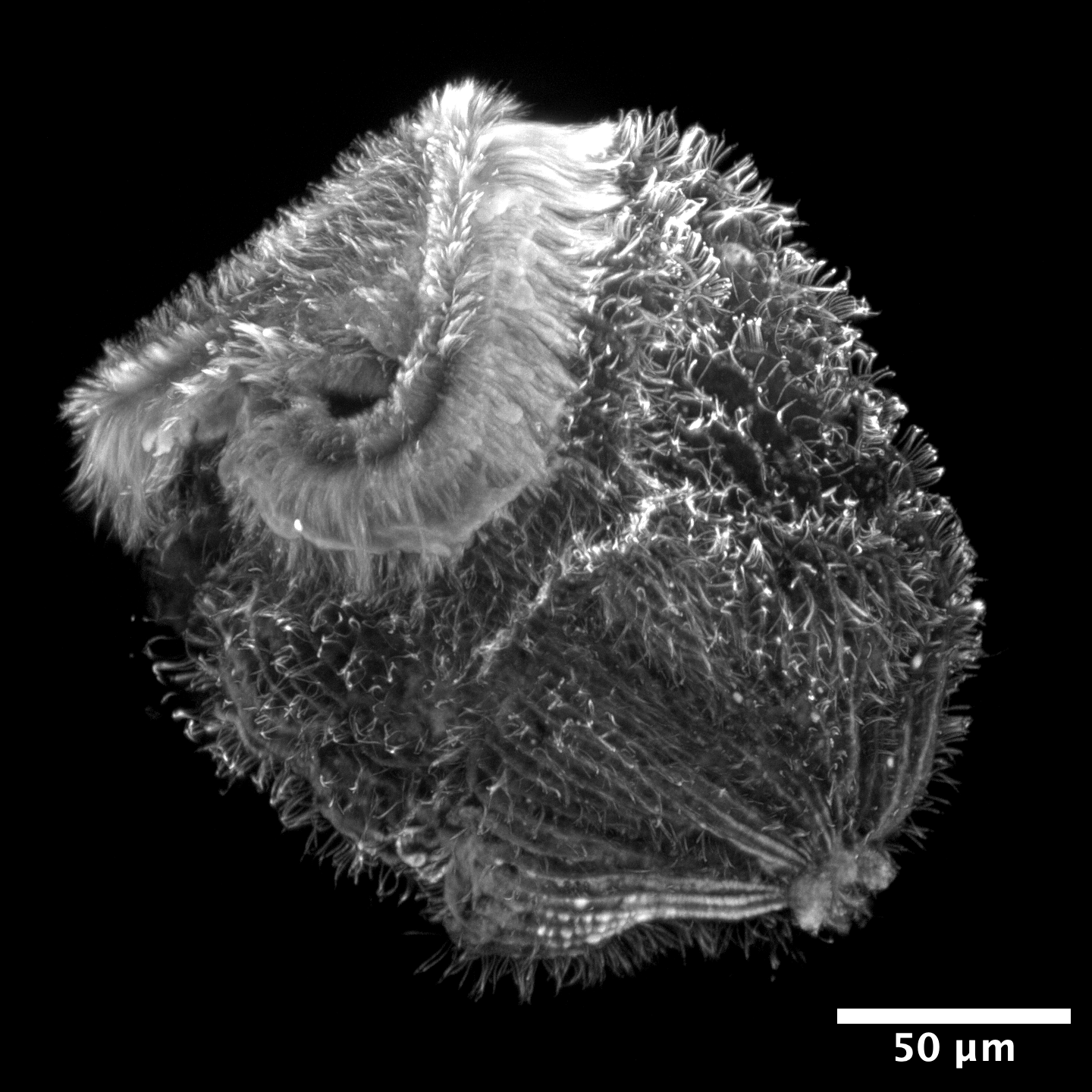
Contributions towards establishing Stentor as a model system for studying cell biology of the nucleus
In addition to my thesis work, I also worked as a part of multiple interdisciplinary teams of scientists and engineers to continue establishing Stentor coeruleus as a model system for cell biology. My work with Stentor contributed to two peer-reviewed publications, of which I am a middle author. While I was a Research Facilitator for the Physiology Course at the Marine Biological Laboratory at Woods Hole, MA, I proposed a project to measure the process of ciliary beat coordination as the oral apparatus of Stentor coeruleus develops during regeneration. As a part of my efforts to establish live imaging protocols for studying the macronucleus of Stentor coeruleus, I became an inventor on a patent with our collaborators from IBM Research.
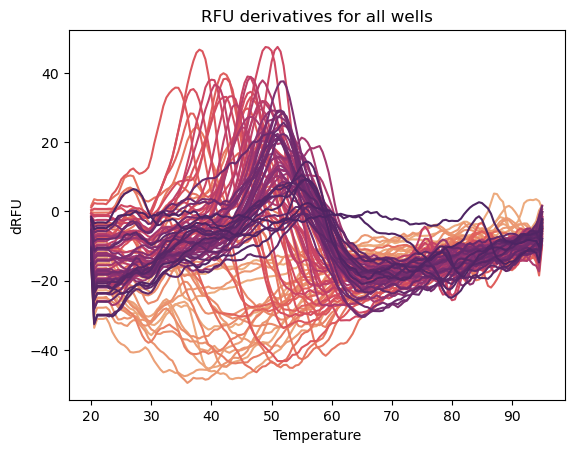
Differential Scanning Fluorimetry Plotting
One of the main challenges of working with purified proteins is knowing which buffer will make the protein “happiest” – i.e. allow the protein to be as stable as possible when it is taken out of a cell and isolated in solution. For commonly studied proteins, oftentimes years of trial-and-error result in buffer recipes that are passed down from mentor to mentee. For a scientist working to purify a new protein, optimizing buffer conditions one by one is tedious and time consuming.
There is a technique called differential scanning fluorimetry that allows the scientist to quantitatively measure their protein’s stability in 96 different buffers simultaneously; turning years of optimization into an experiment that takes an afternoon. Not only does this allow the scientist to choose the one buffer condition that promotes stability the most, but it can also be used to uncover interesting trends in protein stability as buffer conditions vary. Differential scanning fluorimetry works by using a qPCR machine and a dye called Sypro Orange to measure the melting temperature of a purified protein. As the protein unfolds at increasing temperatures, the dye binds to the exposed hydrophobic residues and its fluorescence increases - this fluorescence is measured by the qPCR machine. Comparing this melting temperature across many different buffer conditions allows us to determine which conditions promote protein stability.
This python tool enables scientists to easily import their data and plot melt peaks to determine which condition promotes maximum stability. This tool also can plot protein stability data as a function of a particular buffer condition, like pH, to learn how different chemical conditions affect protein stability trends. To use the tool, download the jupyter notebook at the github link below. Follow the instructions in the notebook to input your buffer conditions and the melt data exported from CFX Maestro software. Then you can plot your melt peaks and create pandas queries to plot data by buffer, pH, salt, etc. For users who are new to using pandas for data manipulation and seaborn for making plots, the notebook provides detailed instructions for every step in the process. The example image above shows a plot containing all of the melt peaks from an entire 96-well plate buffer screen.
Access the github repository here
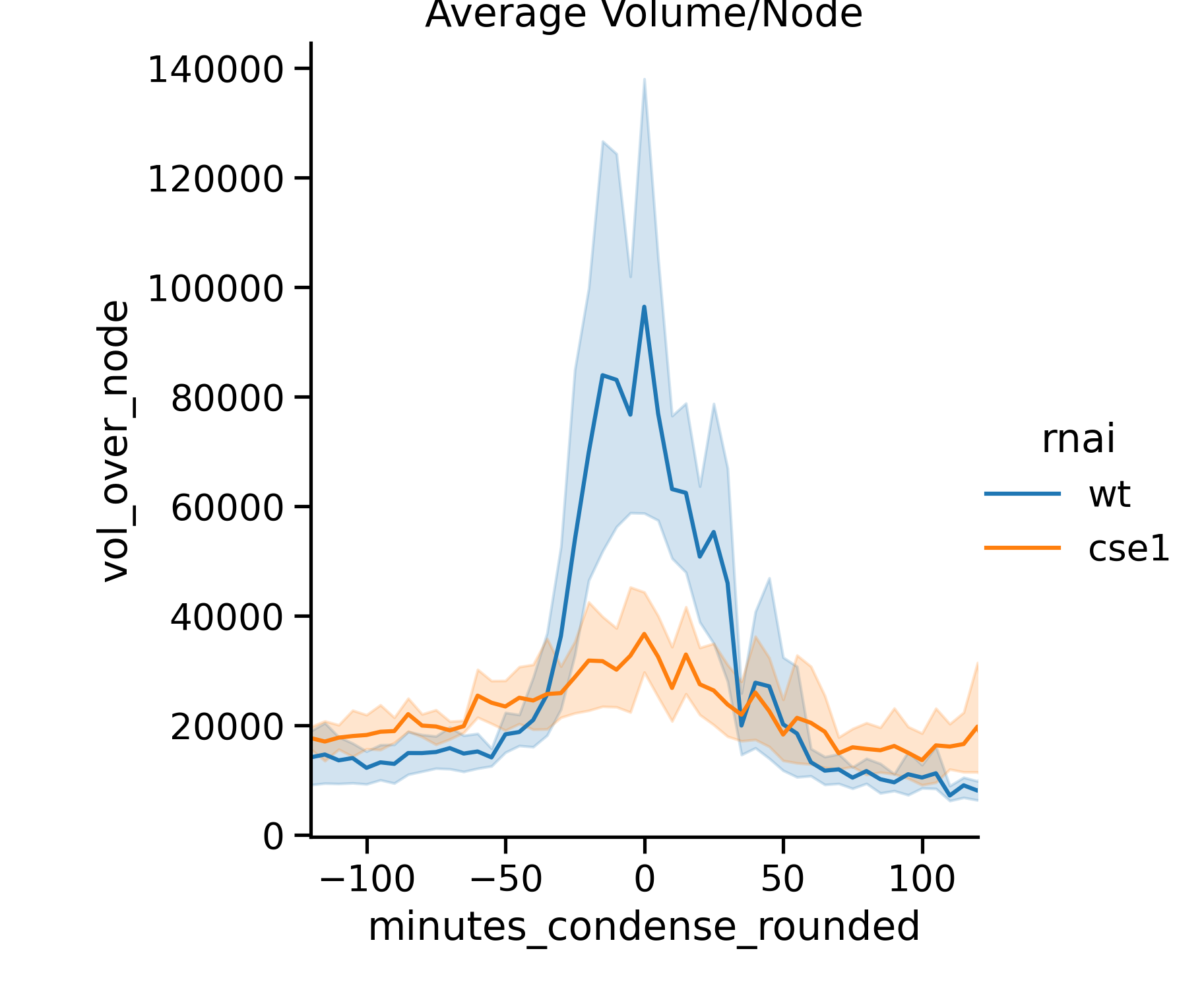
Stentor Macronuclear Volume Analysis
It is very tricky to get 3D images of live Stentor - they are extremely thick samples for microscopy and they are very fast swimmers! One way to calculate the volume of Stentor’s macronucleus is to gently compress the cell in a rotocompressor, enough to slow the cell down but not enough to compress the macronucleus inside. This is a workflow that involves making a mask of the macronucleus, aligning the nodes along an axis of rotation, and then calculating the volume of the entire macronucleus. This was one of my first coding projects that involved jupyter notebooks and image analysis.
Access the github repository here
Code Snippets
In addition to the more complete data analysis workflows above, I’ll also often make smaller scripts to plot data, prep data for analysis pipelines, etc. Feel free browse this repository and use anything that would be helpful for your work!
Access the github repository here
Science and art have always gone hand-in-hand for me. From communicating scientific concepts clearly in graphical form, to making art inspired by science, to drawing to hone my observational skills, my practices in both fields reinforce each other. Click on each image for a closer look.
Scientific Figures
These are vector graphics figures that have been included in scientific publications:
Science-Inspired Art
Work inspired by my colleague's work, my own experiences in research, and the scientific literature:
Observational Work
My more traditional art is inspired by botanical forms. I draw and paint plants I encounter on hikes, discover at botanical gardens, or grow in my container garden.
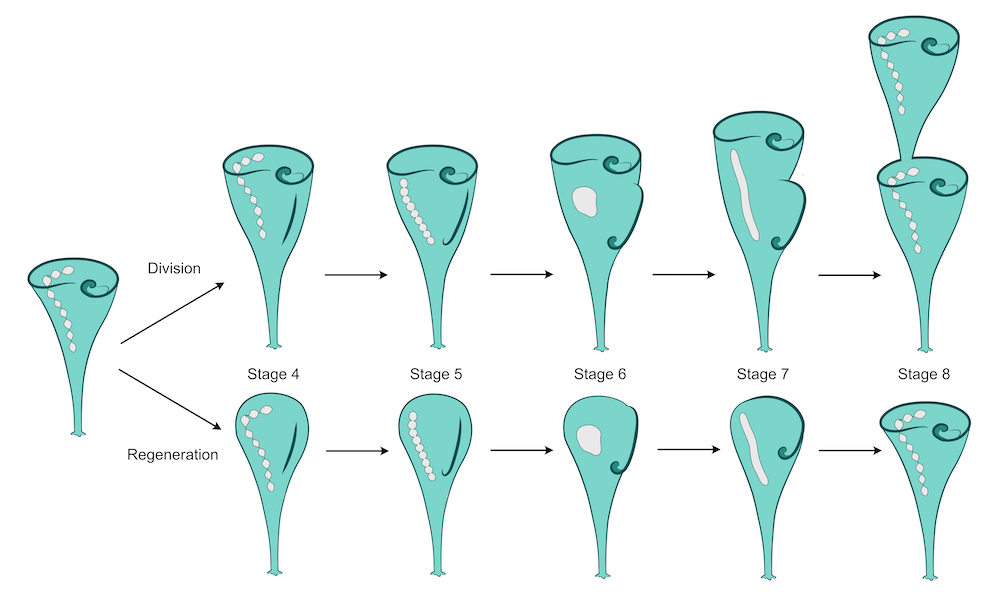
Diagram of the stages of regeneration in Stentor coeruleus. This is a part of my paper on the regulation of nuclear size and shape in Stentor:
McGillivary, R.M., Sood, P., Hammar, K., and Marshall, W.F. (2023). The nuclear transport factor CSE1 drives macronuclear volume increase and macronuclear node coalescence in Stentor coeruleus. iScience, 107318. Read the article here
Back to Art
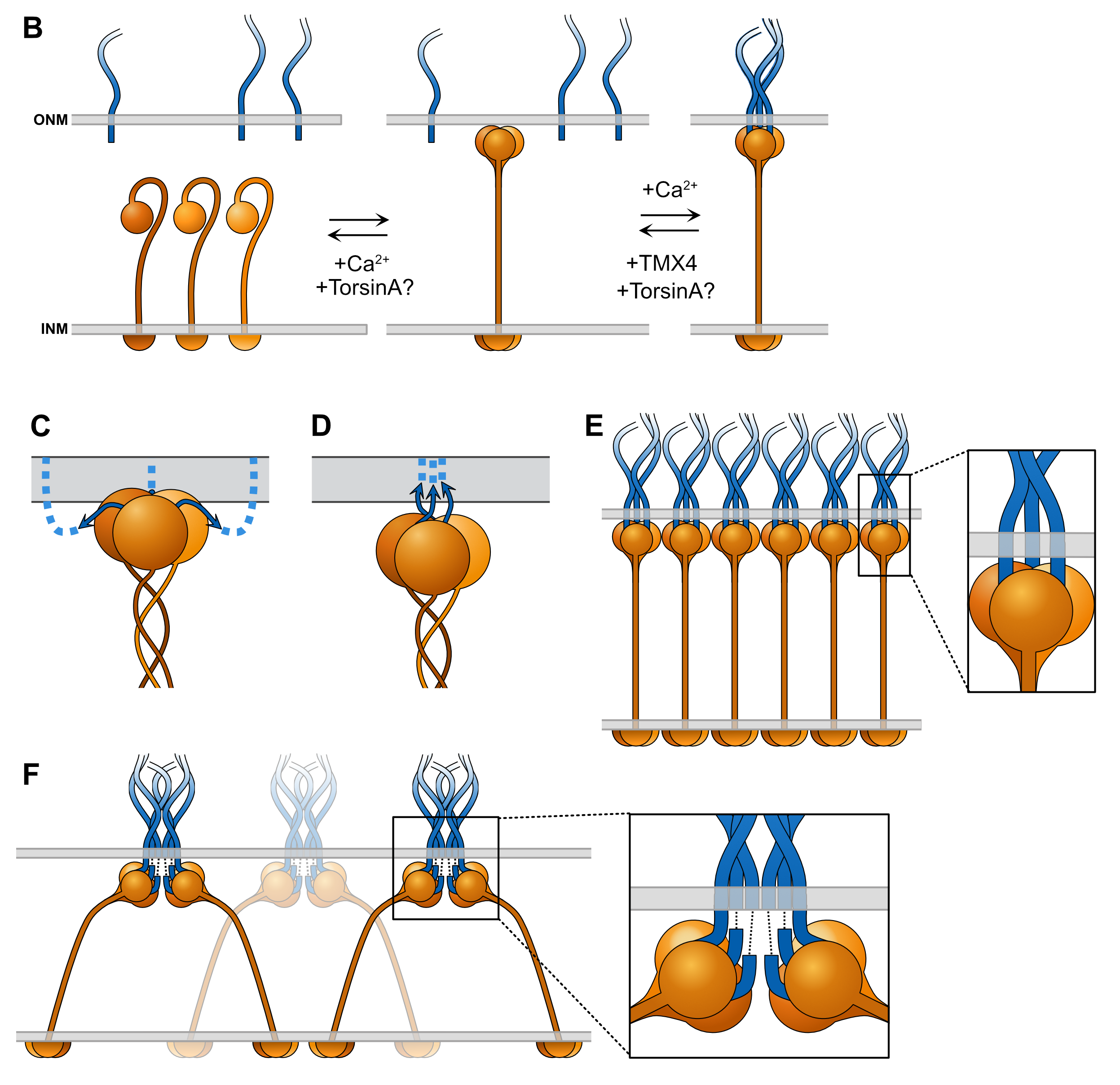
Diagram of various stages and potential configurations for the assembly of the linker of nucleoskeleton and cytoskeleton (LINC) complex. This is a part of my review article:
McGillivary, R.M., Starr, D.A., and Luxton, G.W.G. (2023). Building and breaking mechanical
bridges between the nucleus and cytoskeleton: Regulation of LINC complex assembly and disassembly. Curr. Opin. Cell Biol. 85, 102260. Access free full-text here
Back to Art
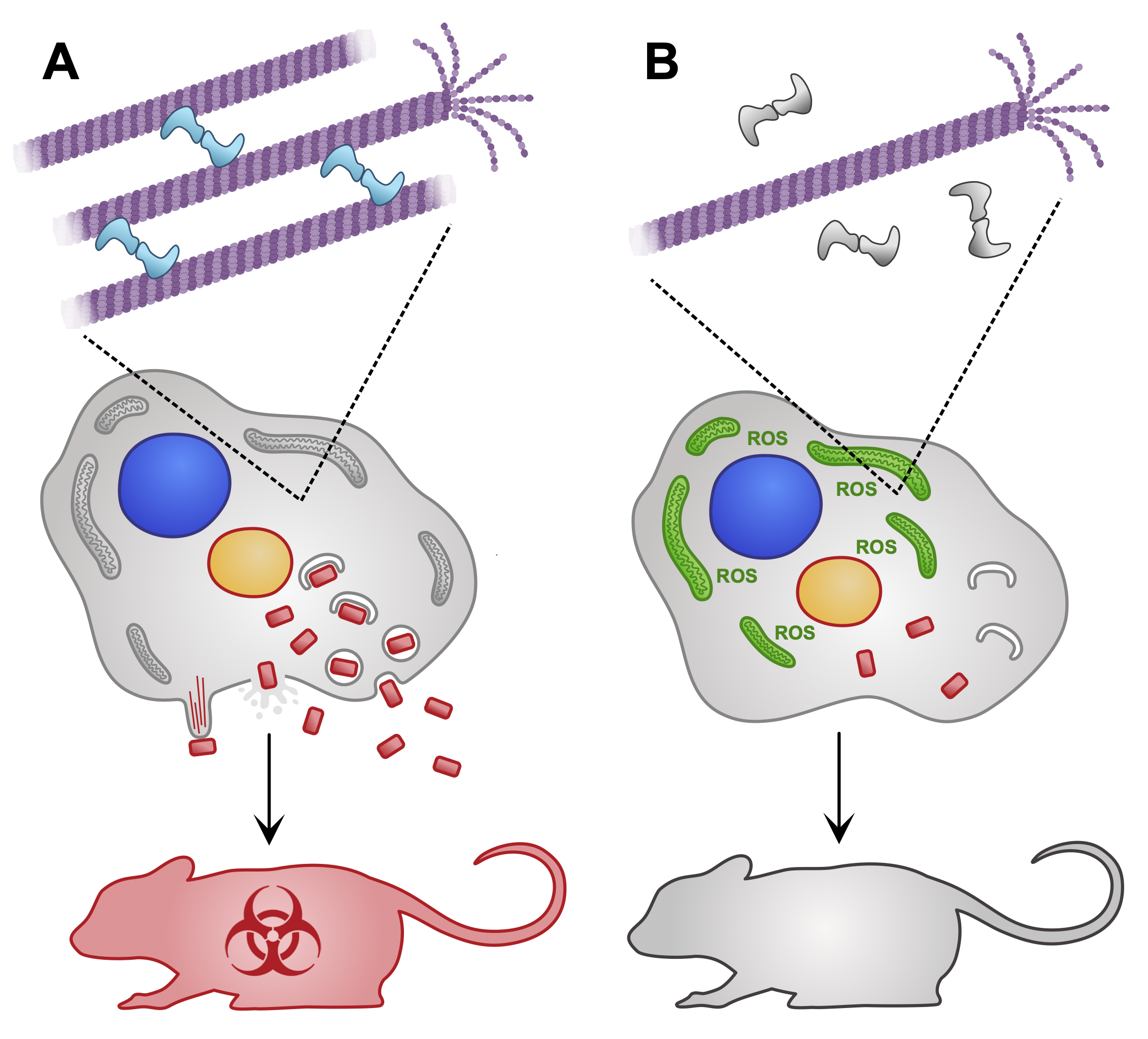
Diagram depicting the poxvirus protein A51R and its role in modulating microtubule stability, and how this has downstream effects that promote the production of more poxviruses. This was for a preview article, where we highlighted and introduced a new research paper that was pubished in the journal by another group.
McGillivary RM, Luxton GWG. Poxvirus A51R: A microtubule maestro and virulence virtuoso. Cell Rep. 2024 Apr 23;43(4):114050. Read the preview article here
Back to Art
Commissioned piece for the Bess Lab at UC Irvine for their website. Their group studies the interactions between small molecules and the microbiome of the gut. Check out their website here!
Back to Art
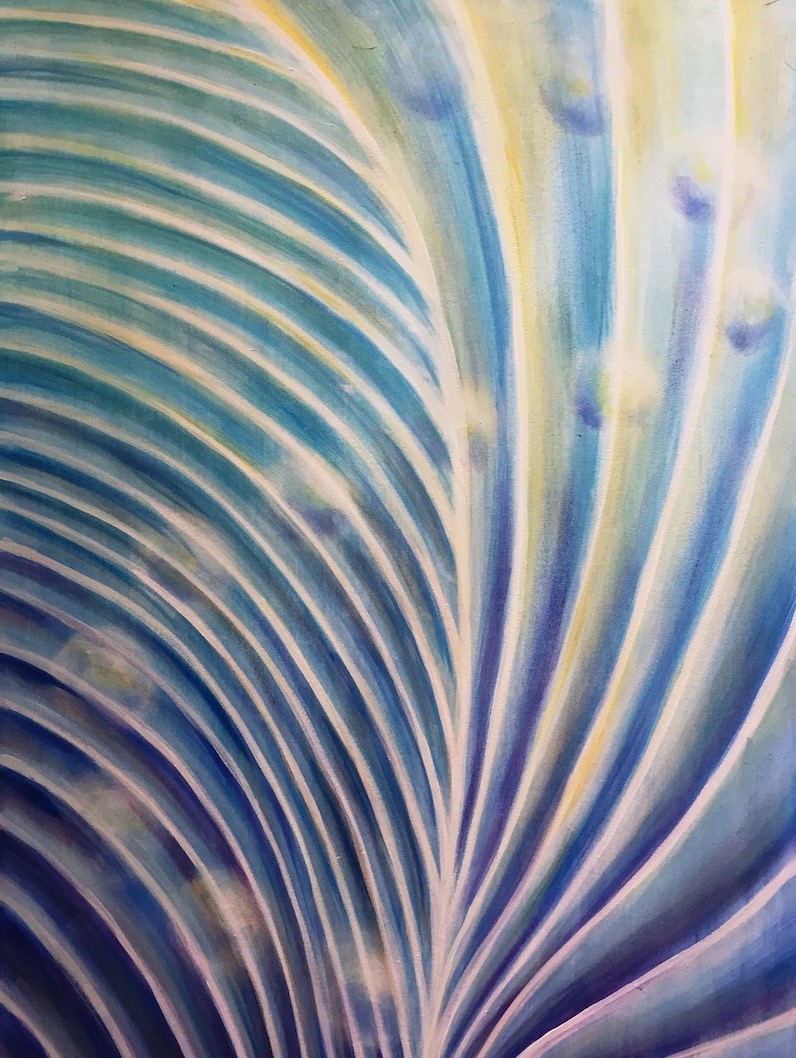

I made this painting just after finishing my qualifying exam in graduate school. I had just spent the past few months thinking non stop about Stentor's macronucleus, and how it is positioned within the cell relative to pattern of blue-green stentorin pigment and clear microtubule bundles on the outside of the cell. Over the following years my research ended up focusing more on the change in shape of the macronucleus instead of how it is positioned. So, the question that inspired this piece has yet to be answered!
Back to Art

In graduate school we spent our first spring quarter taking mini-courses, where we spend two weeks reading and discussing a scientific paper every day. For the evolution mini-course, I decided to also challenge myself to make a drawing every day that was inspired by each paper. This piece was inspired by a paper about the interplay between molecular redundancy and morphological variation.
Back to Art

A quick colored pencil drawing of one of the roses I have been growing in my container garden
Back to Art

An acrylic painting of a barrel cactus
Back to Art

A watercolor and colored pencil drawing of an Echeveria succulent plant
Back to Art
Technical Talks
I am enthusiastic about presenting my work to a wide range of scientific communities, and in venues both large and small. I have presented at five international conferences and have also given additional talks at department seminars, lab retreats, and regional meetings. I have presented to audiences of over one hundred scientists at major conferences, and to smaller focused audiences at lab retreats. Selected talks are listed below:
Outreach

Take Your Own Cell-fie
Cells to Self Exhibition Internship Project
The Exploratorium | San Francisco, CA | 2018 - 2019
I developed a new demonstration by adding a computational spin to an existing demonstration at the Exploratorium that let visitors see their cheek cells under a microscope. Using ImageJ macros, I developed and coded a method to take a photo of the visitor’s face with a regular camera, and computationally merge it with a microscope image of their own cheek cells. This created visually striking images that demonstrated that we are made of trillions of cells (my own cell-fie above is an example of the output!). I would also walk the visitor through the cell images to explain what they were looking at, and answer a wide variety of cell biology questions. I also created training frameworks and conversation flow guides that enabled museum staff to also perform the demonstration successfully. This demonstration was selected to be featured at the opening night of the Cells to Self Exhibition, an After Dark Event, and became a regularly performed demonstration at the museum.
Presenter at Science Festival Exhibits
Marshall Lab at UCSF and NSF Center for Cellular Construction
I volunteered to present demonstrations developed by my colleagues at public science events, interacting with thousands of visitors. These projects featured topics like regeneration, molecular self-assembly, and DIY microscopy - though the exhibit I presented most often was titled “Can Cells Learn?” and featured Stentor cells that could habituate to stop responding to mechanical tapping on their culture dishes.
Community Building
Microscopy Workshop Planning Committee Member
2023 Imaging Workshop for Emerging California Scientists
UC Davis BMCDB Graduate Group + Advanced Imaging Center at Janelia
Davis, CA | 2022 - 2023
I co-organized a microscopy and image analysis workshop for undergraduate and incoming graduate students. I coordinated fundraising efforts among graduate students, faculty, and the UC Davis Foundation and Corporate Engagement Office, personally securing a $1,000 corporate donation. I also engaged with workshop participants to share research experiences and discuss career pathways in microscopy-based research.